#include "embedsom_cuda.h"#include <limits>#include "bitonic.cuh"#include "cooperative_groups.h"#include "cuda_runtime.h"#include "device_launch_parameters.h"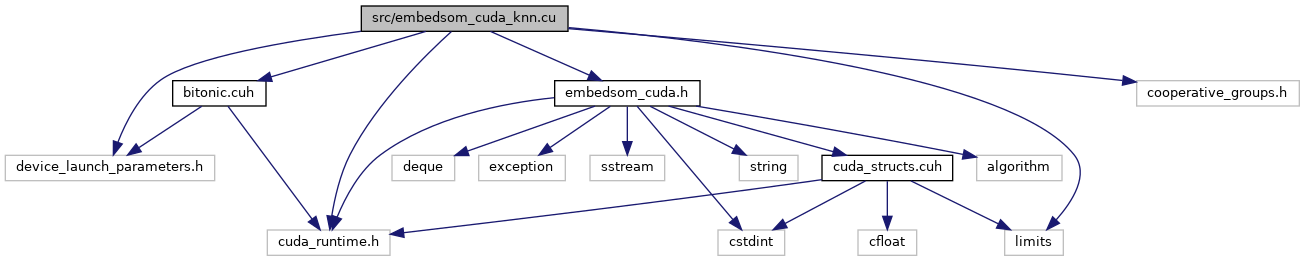
Go to the source code of this file.
Functions | |
| template<typename F = float> | |
| __inline__ __device__ F | sqeucl (const F *__restrict__ lhs, const F *__restrict__ rhs, const uint32_t dim) |
| Compute squared euclidean distance. More... | |
| template<typename F = float> | |
| __inline__ __device__ void | bubbleUp (knn_entry< F > *const __restrict__ topK, uint32_t idx) |
| Bubble a knn-entry up a list, insertsort-style. More... | |
| template<typename F > | |
| __global__ void | topkBaseKernel (const F *__restrict__ points, const F *const __restrict__ grid, knn_entry< F > *__restrict__ topKs, const uint32_t dim, const uint32_t n, const uint32_t gridSize, const uint32_t k) |
| Base kernel for kNN computation. More... | |
| template<typename F , int K> | |
| __global__ void | topkBitonicOptKernel (const F *__restrict__ points, const F *const __restrict__ grid, knn_entry< F > *__restrict__ topKs, const uint32_t dim, const uint32_t n, const uint32_t gridSize, const uint32_t actualK) |
| Bitonic kNN selection. More... | |
| template<typename F , int K = 2> | |
| void | runnerWrapperBitonicOpt (const F *points, const F *grid, knn_entry< F > *topKs, uint32_t dim, uint32_t pointsCount, uint32_t gridSize, uint32_t actualK) |
| Start the bitonic kNN with the right parameters. More... | |
Function Documentation
◆ bubbleUp()
| __inline__ __device__ void bubbleUp | ( | knn_entry< F > *const __restrict__ | topK, |
| uint32_t | idx | ||
| ) |
Bubble a knn-entry up a list, insertsort-style.
Definition at line 55 of file embedsom_cuda_knn.cu.

◆ runnerWrapperBitonicOpt()
| void runnerWrapperBitonicOpt | ( | const F * | points, |
| const F * | grid, | ||
| knn_entry< F > * | topKs, | ||
| uint32_t | dim, | ||
| uint32_t | pointsCount, | ||
| uint32_t | gridSize, | ||
| uint32_t | actualK | ||
| ) |
Start the bitonic kNN with the right parameters.
This function recursively scans (in compiler!) for a sufficient K to contain the actual number of neighbors required and runs the correct instantiation of topkBitonicOptKernel().
Definition at line 199 of file embedsom_cuda_knn.cu.
◆ sqeucl()
| __inline__ __device__ F sqeucl | ( | const F *__restrict__ | lhs, |
| const F *__restrict__ | rhs, | ||
| const uint32_t | dim | ||
| ) |
Compute squared euclidean distance.
Definition at line 42 of file embedsom_cuda_knn.cu.
◆ topkBaseKernel()
| __global__ void topkBaseKernel | ( | const F *__restrict__ | points, |
| const F *const __restrict__ | grid, | ||
| knn_entry< F > *__restrict__ | topKs, | ||
| const uint32_t | dim, | ||
| const uint32_t | n, | ||
| const uint32_t | gridSize, | ||
| const uint32_t | k | ||
| ) |
Base kernel for kNN computation.
Each thread iterates over whole point grid and computes kNN for a specified point using insertion sort in global memory.
Definition at line 72 of file embedsom_cuda_knn.cu.
◆ topkBitonicOptKernel()
| __global__ void topkBitonicOptKernel | ( | const F *__restrict__ | points, |
| const F *const __restrict__ | grid, | ||
| knn_entry< F > *__restrict__ | topKs, | ||
| const uint32_t | dim, | ||
| const uint32_t | n, | ||
| const uint32_t | gridSize, | ||
| const uint32_t | actualK | ||
| ) |
Bitonic kNN selection.
This uses a bitonic top-k selection (modified bitonic sort) to run the kNN search. No inputs are explicitly cached, shm is used for intermediate topk results
- Template Parameters
-
K is number of top-k selected items and number of threads working cooperatively (keeping 2xK intermediate result in shm) note that K is a power of 2 which is the nearest greater or equal value to actualK
Definition at line 122 of file embedsom_cuda_knn.cu.